We have a big technical design need.
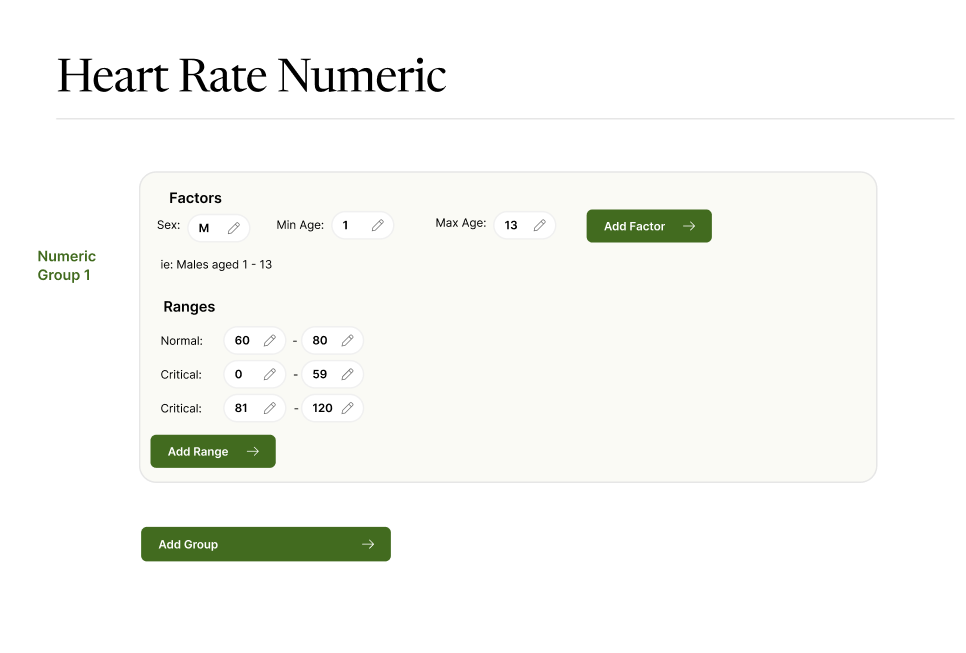
For OpenMRS to be useful in general OPDs, we must have age-based-ranges for things like Vital Signs and Labs.
Why: Because a patient of any age can come through the door - from 1 hour old to 100+ years old. And many Normal vs Abnormal ranges, especially Vital Signs, are different depending on Age (but not just age - also Sex and other variables, but let’s focus on Age for now).
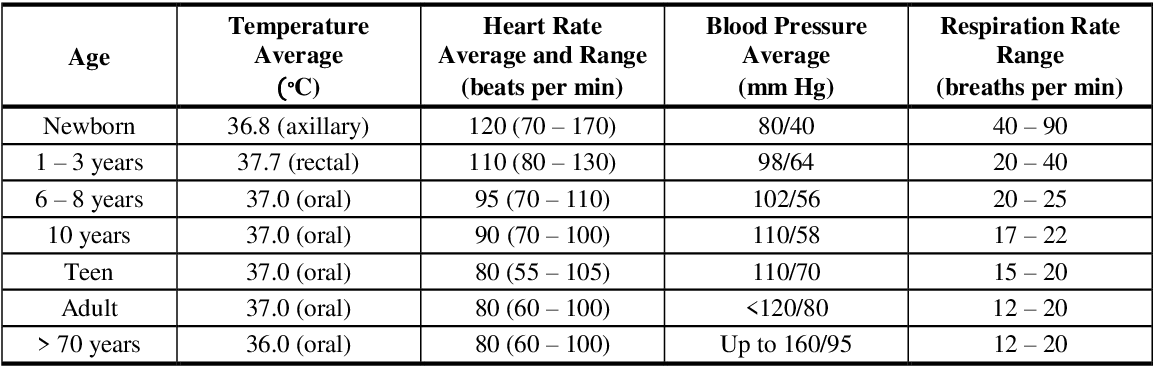
Here are some examples of just how different these ranges can be. Note how extremely different the normal range is for a newborn vs an adult: Something normal for an adult is in fatal-range for a newborn, and vice-versa!
(Source)Lab Ranges Example
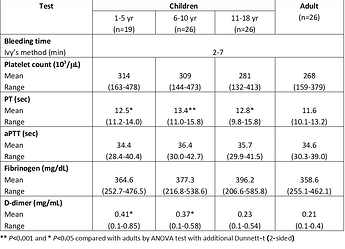
The same is true for many lab tests:
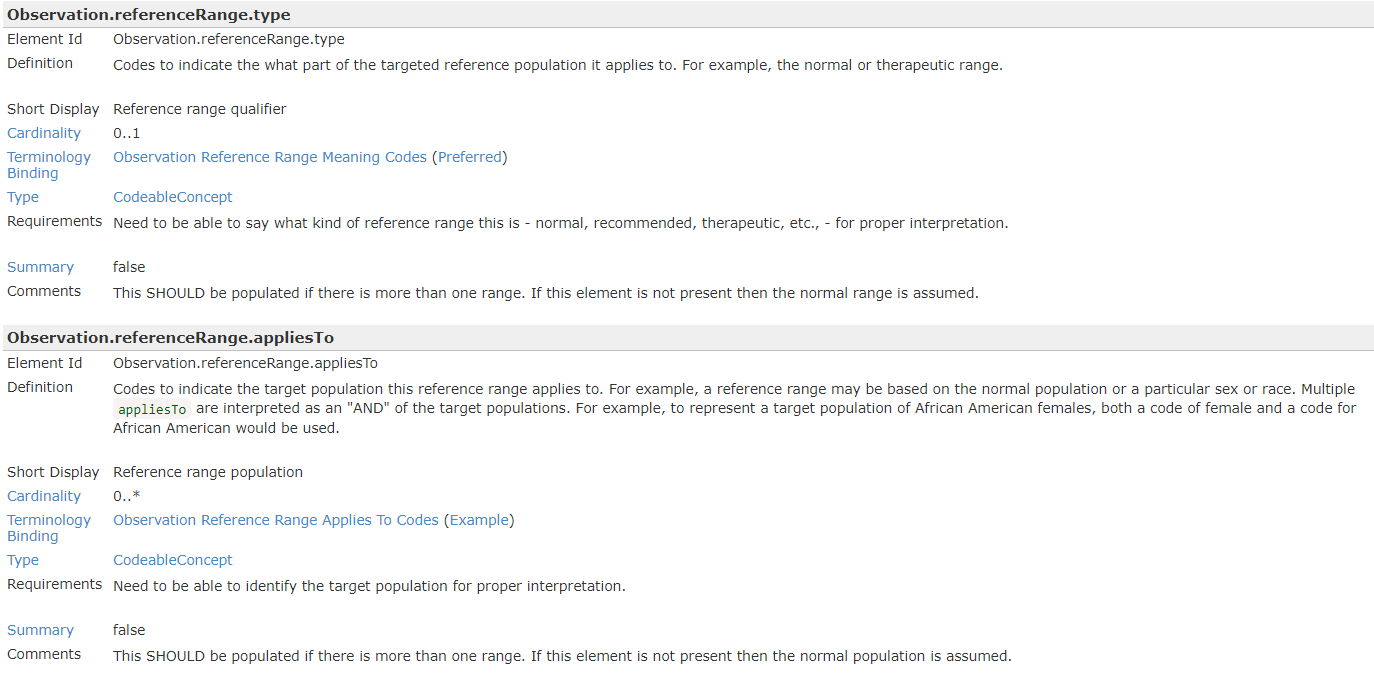
(Source)Sex-Based and other Normal Range Differences
While ages are the most urgent use case for O3 for OPD right now, it’s not just ages that matter. Other variables about a person can change what’s a normal range for them, like kidney function, etc. But the most common example after ages is Sex. See these examples where what’s normal for a man is abnormal for a woman, and vice-versa, as this Thyroid Treatment example shows:
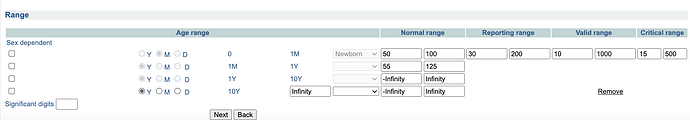
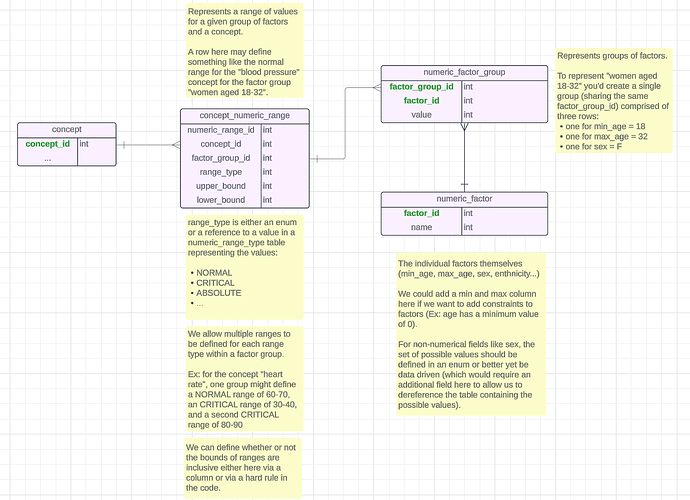
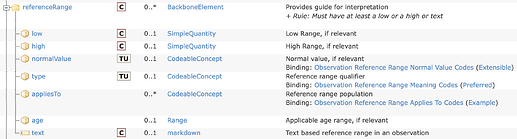
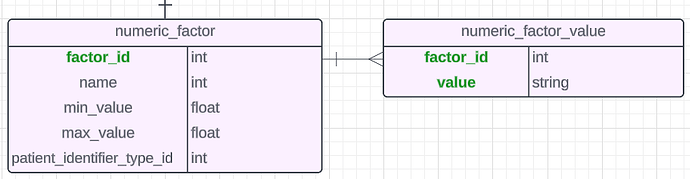
(Source)Currently our normal/abnormal result ranges are metadata directly on concepts, meaning, you can only have 1 range per variable.
@dkayiwa @burke @ibacher Can you start thinking about this? We need a plan, and as a community we need to implement this within the next 2-3 O3 releases or the next platform release. Because this is needed NOW, in the field.